Anthurium, belonging to the Araceae family is a crop of commercial importance in India and many other countries. It is cultivated for its beautiful cut flowers, cut foliage and as a potted plant. In 2016, anthurium grown in greenhouses on the RHREC campus, GKVK Bengaluru, Karnataka, India, showed various types of disease symptoms with an initial incidence of approximately 25%. In 2018, the disease incidence had increased to 80% along with heavy infestations of thrips. Plants showed chlorotic and necrotic lesions on spathes (Fig. 1) and leaves (Fig. 2), and malformation of spathes and necrotic streaks on flower stalks. These symptoms were observed on anthurium cvs. Acropolish, Cheers, Fire, Tropical and Xavia. The symptoms resembled those caused by Tomato spotted wilt virus on anthurium reported from Hawaii (Uchida et al., 1999).
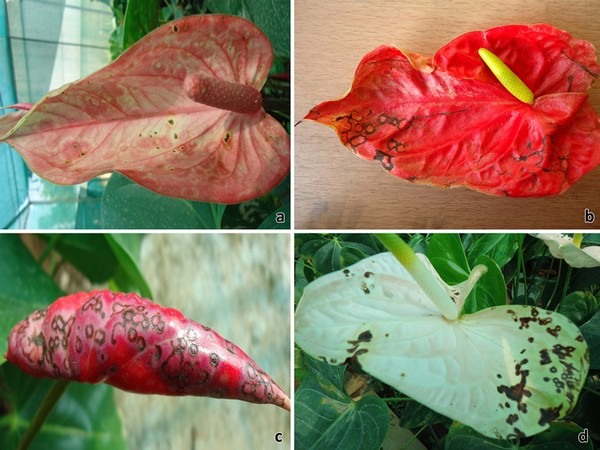

Figure 1: Symptoms of Groundnut bud necrosis virus infection on anthurium spathes in cultivars: a) Xavia, b) Tropical, c) Fire and d) Acropolish.
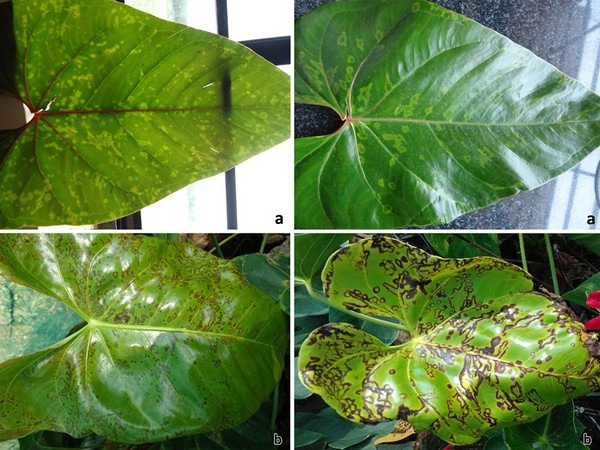

Figure 2: Symptoms of Groundnut bud necrosis virus infection on anthurium leaves: a) chlorotic lesions, and b) necrotic lesions.
Twenty samples comprising either leaves or spathe were collected randomly from symptomatic plants and tested along with buffer and healthy plant controls for the presence of serogroup IV tospoviruses using polyclonal antibodies raised against Groundnut bud necrosis virus (GBNV) (reported to detect all serogroup IV orthotospoviruses (Mandal & Jain, 2010)) by DAC-ELISA. ELISA results were positive for all symptomatic plants with mean absorbance values (A405) ranging from 0.65 to 1.4 compared to 0.12 in the healthy plant control. To confirm the presence of an orthotospovirus species, total RNA was extracted from symptomatic and healthy leaf samples using the SpectrumTM Plant Total RNA Kit (Sigma-Aldrich USA) and subjected to RT-PCR analysis. The serogroup IV orthotospovirus group-specific reverse primer (5′-AGAGCAATCGAGGCGCTAATAA-3′) and individual species-specific forward primers designed using sequence analysis (BLAST or CLUSTAL O (1.2.4) and Oligo Analyzer (1.0.2) were used for RT-PCR. An amplicon of the expected size (898 bp) was obtained with the GBNV-specific forward primer (5′-TTACAATTCCAGCGAAGGAC-3′) from all symptomatic plants and but not from healthy samples (Fig. 3). The product was cloned and sequenced and the sequence deposited in GenBank (Accession No. MF989441). Analysis of the obtained sequence revealed that it had 89.18-98.07% nucleotide and 87.32-98.91% amino acid identity with the N gene of GBNV isolates deposited in GenBank from different crops and regions. The highest identity was observed with a GBNV tomato isolate from Coimbatore (AY472081). Phylogenetic analysis by the neighbour-joining method using MEGA 7 software indicated that it grouped with other GBNV isolates (Fig. 4).
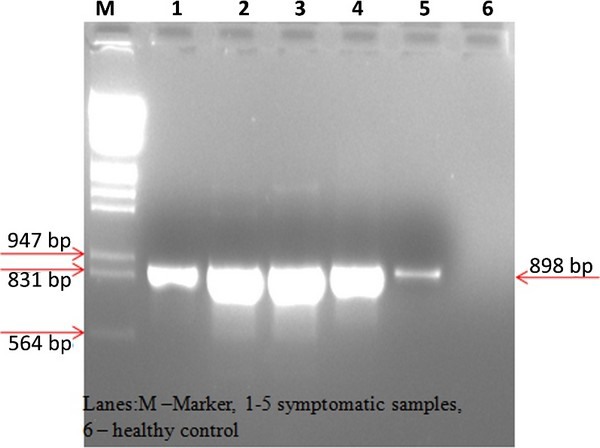
Figure 3: Agarose gel analysis of RT-PCR products amplified using serogroup IV tospovirus-specific reverse and Groundnut bud necrosis virus-specific forward primers.

Figure 4: Phylogenetic analysis of N gene sequences of Groundnut bud necrosis virus (GBNV) on anthurium with other isolates of GBNV and Tomato spotted wilt virus.
The occurrence of members of the genus Orthotospovirus, such as Tomato spotted wilt virus and Impatiens necrotic spot virus, which are transmitted by several species of thrips, on anthurium has been reported from different parts of the world (Uchida et al., 1999, Ghotbi & Shahraeen, 2012). In India, GBNV is the most widely distributed member of the genus Orthotospovirus and infects a wide variety of plant species (Mandal et al., 2012). However, to the best of our knowledge, anthurium is a new host for GBNV, not only in India but also worldwide. The severity of symptoms observed on anthurium suggests that, GBNV is an important and emerging virus on this ornamental plant with the potential to cause economic damage and therefore merits attention.










